Hit Identification at NovaData
The timely and effective screening of chemical databases for putative bioactive molecules is an important initial phase in many research projects. NovaData provides a tailored approach to identify solutions for this challenge with an industry standard virtual screening platform which is adaptable to allow for the design of bespoke chemical libraries based on your chemistry or from commercial sources.
Fragment-based approaches
Fragment-based drug discovery is a powerful means to identify novel starting points for discovery projects. Virtual screening fragment libraries is a cost effect means to mine the extensive fragment space to identify novel fragment binding modes against a known biological target. However, designing the optimal fragment library for the desired biophysical or biochemical screening method can be challenging. At NovaData, we have the experience in designing bespoke fragment libraries and we have the capabilities to identify fragments using a combination of Docking and Quantum Mechanical approaches.
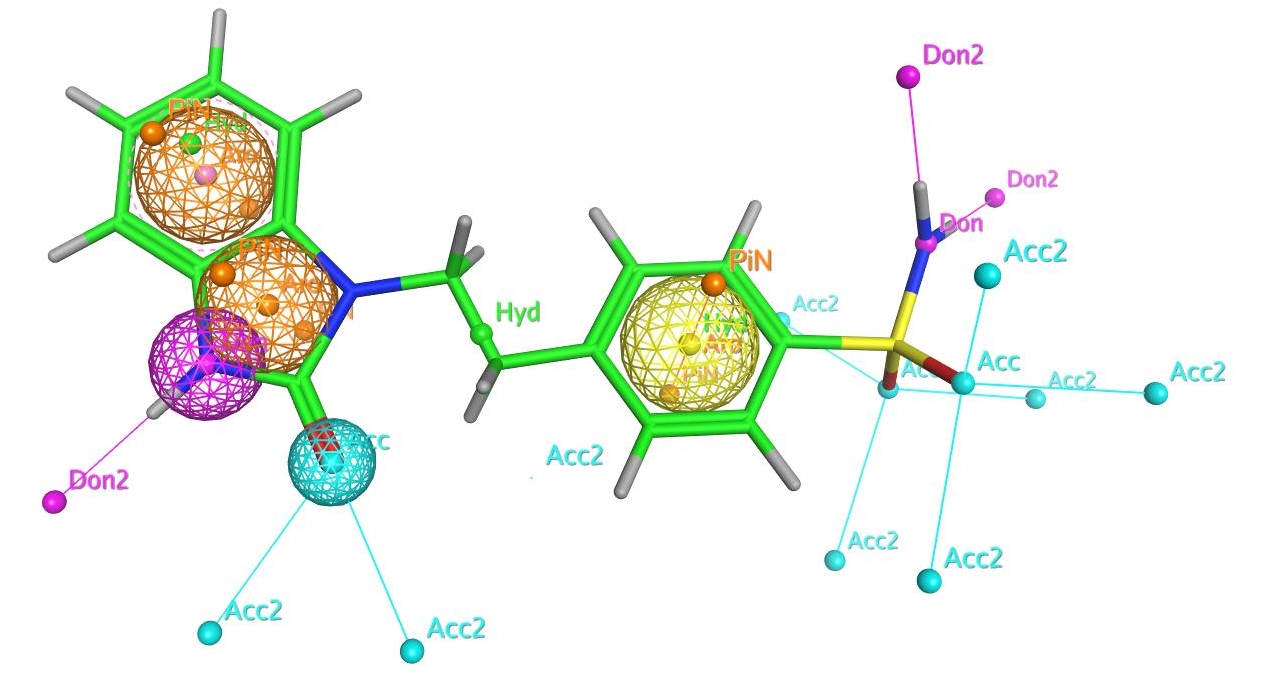
Ligand-based approaches
At NovaData we screen chemical databases using the latest technology in 2D and 3D ligand-based approaches, including:
- Molecular fingerprints and 2D/3D property-based descriptors
- Pharmacophores
- ROCS - shape and electrostatic based molecular alignment
- QSAR and predictive modelling
Our approach is to perform a thorough search using a combination of approaches, and where appropriate compliment the search with QSAR model generation. Candidate molecules are then filtered based on a number of properties to meet your project hit criteria. Hits can be prioritized on a number of parameters including off-target selectivity, chemical diversity, property space, ease of synthesis and commercial cost and availability.
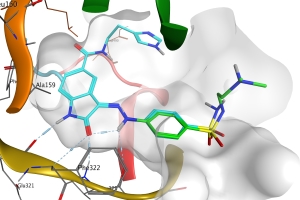
Structure-based approaches
There are many projects which have identified a biological target but have no chemical starting point for hit identification. This presents an opportunity to perform structure-based design. More often a hit is known and the challenge is to find new chemical matter with the same mode of target engagement or an alternative mode of binding. At NovaData we have extensive experience in developing 3D structure models to enable structure-based design, either from X-ray, EM, NMR data or from homology modelling. We utilise high throughput docking and target-based pharmacophore modelling to screen chemical databases. And whenever possible we also perform detailed analysis of ligand-target molecular interactions to aid in rational design and candidate molecule prioritization. The challenge of target selectivity can also be addressed through off-target structure-based analysis.
Our approach is to perform structure-based approaches in combination with ligand-based methods, giving clients a wealth of valuable information to increase the success in early discovery research.
